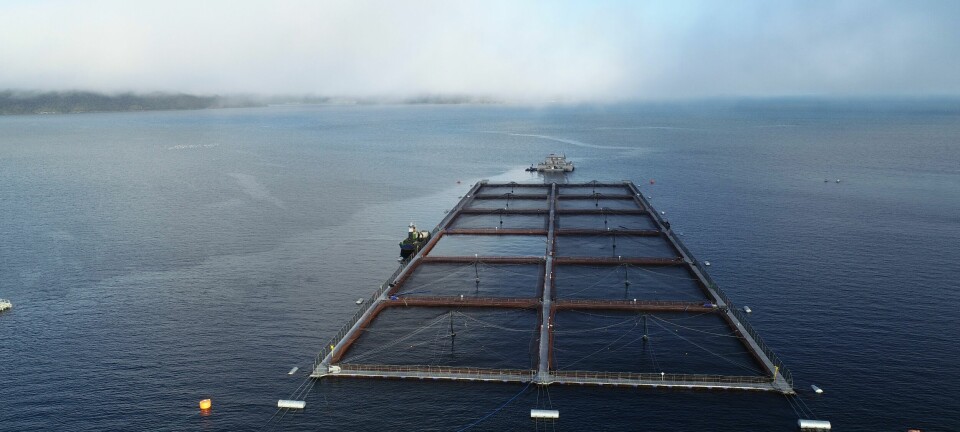
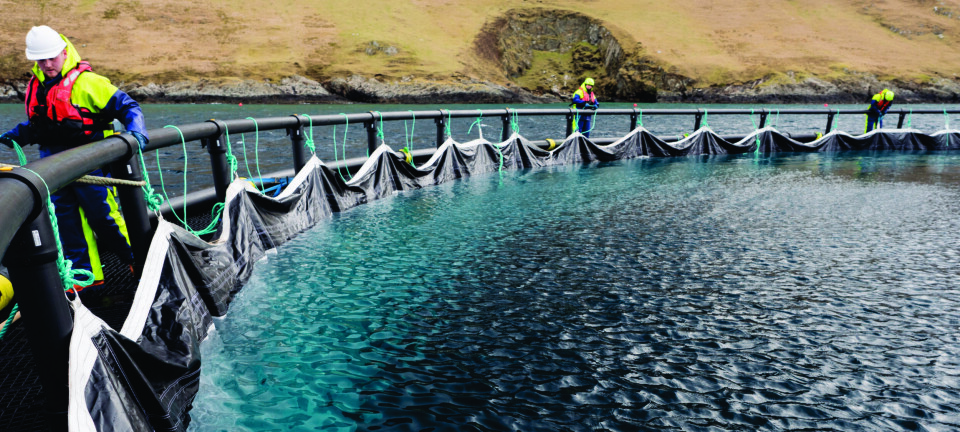
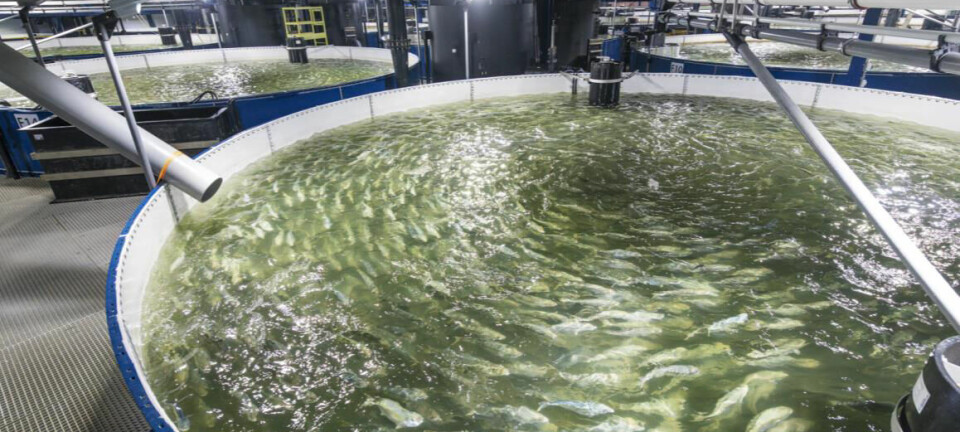
Aquaculture lagging behind in selective breeding
There is huge potential to improve disease resistance in aquaculture worldwide if more fish farmers adopt selective breeding, an Edinburgh fish genetics expert has said.
Professor Ross Houston pointed out that less than 10 per cent of aquaculture production currently derives from selectively bred stocks, lagging significantly behind terrestrial animal and plant farming.
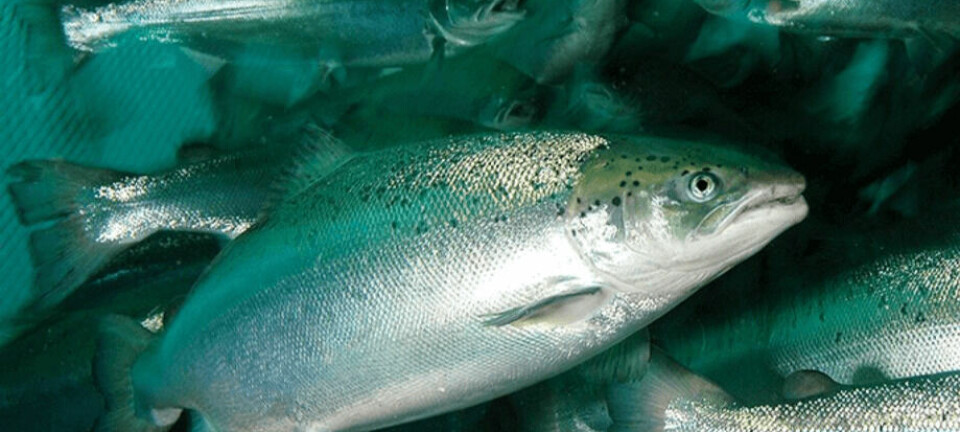
In a review for the Brazilian Journal of Animal Science, titled Future directions in breeding for disease resistance in aquaculture species, the Roslin Institute's Chair of Aquacultural Genetics writes: “Infectious disease is a major constraint for all species produced via aquaculture. The majority of farmed fish and shellfish production is based on stocks with limited or no selective breeding. Since disease resistance is almost universally heritable, there is huge potential to select for improved resistance to key diseases.”

Houston points out that while there are vaccines and medicines available for several farmed aquatic species, particularly finfish, which aid in the prevention and control of disease, they are often expensive and only partially effective.
Evolution of resistance
He adds that blanket treatments are often used (e.g. in feed), which can lead to evolution of resistance in the pathogen, an example being “the emergence of drug-resistant strains of ectoparasitic copepod sea lice, due to extensive use of medicines (Aaen et al., 2015). Therefore, a major and increasingly important component of disease control is selective breeding to produce stock with improved resistance to key pathogens, exploiting naturally occurring genetic variation (heritability) for resistance in farmed aquaculture populations.”
Houston concludes: “Disease resistance is almost universally heritable and is a key goal of existing selective breeding schemes. Several success stories of mass selection, family selection, and marker-assisted selection are evident. Gathering appropriate phenotypes from disease challenge or field experiments is pertinent for making genetic progress. Genomic selection is the state-of-the-art for modern aquaculture breeding schemes and offers substantial improvements in selection accuracy over pedigree-based methods.
Major challenge
“Driving down the cost of genomic selection, and specifically generation of genome-wide genetic marker datasets, is a major current goal. Genotype imputation and low-cost genotyping by sequencing are likely to contribute.
“Predicting disease resistance of distantly related animals to those with trait records is a major future challenge, which is directly related to the identification of causative variants. Genome editing technology is likely to play a key role in identifying causative variation and has potential for breeding disease-resistant animals in aquaculture.”
Houston's research group discovered a major QTL affecting resistance to Infectious Pancreatic Necrosis (IPN) virus which has been extensively applied in the aquaculture industry. He has also been involved in the development of tools to investigate and improve resistance to Oyster Herpes Virus.